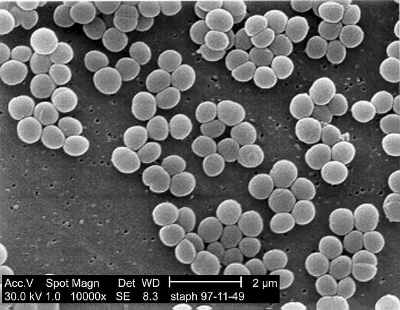
The human skin is the body’s first line of defense against pathogens that your body comes in contact with. Just like the gut and mouth, the skin lives in communion with bacteria. One important unanswered question that many scientists have is why commensal bacteria do not trigger an inflammatory immune response when they come in contact with the skin. An article published by Cell Press explores exactly this question, looking specifically at regulatory T cells (treg, a type of white blood cell that plays a major role in establishing homeostasis of the immune system).
Researchers engineered the genes of Staphylococcus epidermidis to produce a specific protein antigen that can be fluorescently viewed. To test whether the immune system plays a role in tolerance of skin commensal bacteria the researchers colonized the skin of 6-week-old mice with this fluorescent protein. Three weeks later the mice were compared to a group of control mice and it was found that pre-colonization with the protein was not enough to establish immune tolerance of the bacterial antigens.
The researchers were curious as to what affect this bacterial antigen on the skin of infant mice had, so the same experiment was done with 7-day-old mice. After 3-4 weeks, when the mice were adult, a significantly diminished immune response to the commensal bacteria could be seen. This shows that exposure during the neonatal period promotes tolerance to commensal bacteria.
After examining adult vs. neonatal skin, this study concludes that there is a difference between the two in terms of immune response and windows of tolerance build-up. Specifically, the period of neonatal skin development seems to be essential in mice for the immune tolerance of commensal bacteria. The implications of this study are important for understanding of the human immune system and bacteria tolerance. Because the skin is our body’s first defense system, it is important to have an understanding as to what mediates its immune response.